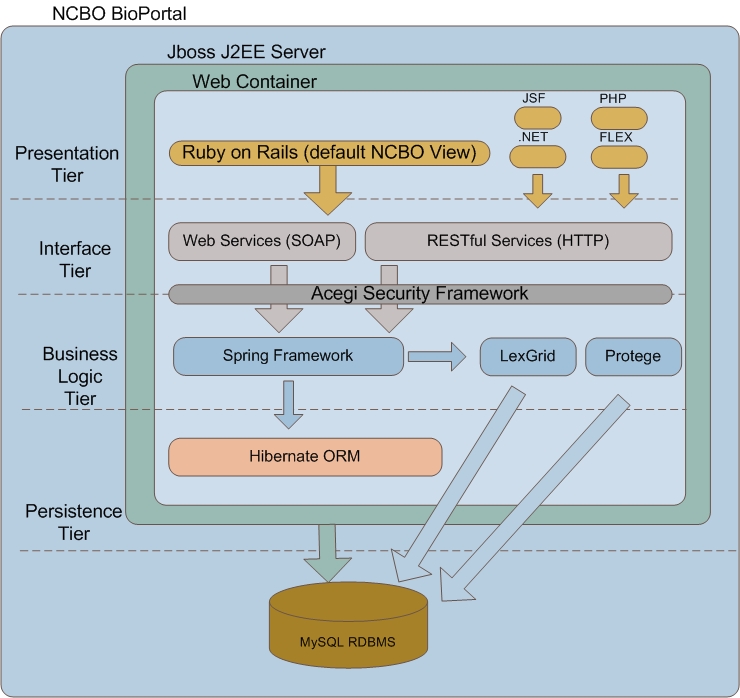
NCBO-OOR Architecture
The NCBO-OOR architecture (essentially the NCBO BioPortal architecture) uses a classic architecture layered approach, which decouples the logic and domain object models between each layer. This approach decouples the versioning and changes in one layer from another. Thus, as software modules evolve (which they always do), the impact on the rest of the software sub-systems is significantly reduced. Furthermore, the BioPortal enforces de-coupling through web services (e.g., REST and SOAP) and the enterprise pattern of “Dependency Injection.” Note that Protégé 3.x has been incorporated into business logic layer (in addition to pre-existing LexGrid) to enable full support of OWL ontologies and the capability to store ontology based meta data, mappings, and marginal notes. The following diagram presents a simple overview of the layers in the BioPortal architecture followed by a description of each.
The Presentation Tier delivers the BioPortal user-interface which currently uses the Ruby on Rails technology. Ruby on Rails is a leading mature UI framework supported by large software development communities. It enables rapid prototyping as well as solid integration with web services.
The Interface Tier consists of both REST and SOAP web services that present all BioPortal capabilities to the upper tiers (e.g., upload ontology, download ontology, display concept, administrative functions, etc). The Presentation Tier is primarly driven by the REST services (currently implemented with RESTlet libraries). Thus, the BioPortal Presentation Tier is just one vanilla consumer of the BioPortal REST services. In the principle of Service-Oriented Architecture, any number of partner developers could consume the set of existing REST services for any number of purposes. For example, one could implement a completely different UI than that currently exposed by the BioPortal. Or, one could simply want to consume a single REST services for integrating into a back-end workflow with no UI at all. Please see NCBO REST Services documentation to learn about consuming the BioPortal REST services.
The Business Logic Tier uses the Spring technology which enables a partner to insert any software implementation that abides to the NCBO-defined interfaces. This is achieved through the “Dependency Injection” enterprise pattern which is core to the Spring framework. For example, if NCI requires use of a different flavor of LexGrid, a module could be easily implemented by NCI using the NCBO interfaces and deployed without having to modify the core BioPortal software. This architectural approach enables incorporation of new capability into the BioPortal through packaging and configuration. The need to do a major BioPortal software release to support a customized implementation is significantly reduced. NCBO-OOR Server-Side Customization provides further details how such a customization can occur.
The Persistence Tier uses the Hibernate technology as a basic object-relational mapping to the back-end relational database. Hibernate is used for storing administrative (e.g., user information) and external ontology data (e.g., ontology attributes specified at upload time). All ontology content is stored in Protege and LexGrid as shown in the Business Logic layer.
Consolidating the Bioportal back-end
While architecturally it is preferable to have a single ontology/terminology back-end repository to reduce complexity, no single repository meets all NCBO requirements. The NCBO must support the OWL language and RDF/XML, OBO, RDF and RRF formats. As shown in the diagram, Protégé 3.x has been incorporated into the business-logic layer to enable full support of OWL ontologies and the capability to store ontology based meta-data, mappings, and marginal notes. LexGrid continues to support the diverse terminology formats required by NCBO (e.g., the RRF format used for UMLS is particularly critical). As shown in the above diagram, the combination of LexGrid and Protégé jointly provide sufficient support for the diverse standards and formats. All OWL ontology requests (e.g., get concept, find concept, and upload) will be routed to Protégé. All other ontology/terminology requests such as RRF and OBO will be routed to LexGrid. Long-term, the NCBO plans to rearchitect the BioPortal to leverage both normalized store and native OWL/RDF triple store in collaboration between Stanford University and Mayo Clinic.
With this in mind, adding, removing, and changing back-ends to the BioPortal is modular by design. The server-side architecture implicitly is based on the principle of aspect-oriented programming. Thus, the majority of server-side component in the BioPortal can be modified to meet the needs of OOR.