NCBO-OOR Server-Side Customization
Introduction
OOR provides server-side flexibility by leveraging the enterprise pattern of "Dependency Injection." This is accomplished by applying the Spring technology which enables a partner to insert any software implementation that abides to the NCBO-defined interfaces purely by configuration. In other words, a different implementation can be deployed without a recompile or rebuild of the entire OOR code-base.
This first OOR implemntation is based on the BioPortal architecture and code-base. Thus, the majority of the developer considerations for NCBO-OOR is generally applicable to the BioPortal (and vice-versa). The author will note differences where appropriate.
BioPortal Server-Side Pattern: Dependency Injection
The gist of dependency injection is that a seperate object, an assembler, populates the implementation of a defined interface that can be used by any consuming object that understands the interface. Spring implements a specific variation of this pattern called "Setter Injection." A significant number of BioPortal server-side capabilities heavily leverage this pattern. This approach enables these BioPortal capabilities to easily swap out different implementations. To learn more about this pattern in general, please see Martin Fowler's excellent exposition on this topic (http://martinfowler.com/articles/injection.html). The following sections elucidate a couple BioPortal-specific examples.
A Simple Example
This section walks-through a simple example where BioPortal leverages the Setter Injection model in the BioPortal Interface/Service Layer. In this scenario, "ConceptRestlet" accepts REST service invocation and delgates concept requests to an internal class named "ConceptServiceImpl." "ConceptRestlet" receives a hook to "ConceptServiceImpl" by having it injected by the Spring framework. "ConceptRestlet" is only aware of the "ConceptService" interface and has no direct reference to "ConceptServiceImpl." Note that for clarity, we will not delve into the REST framework (i.e., Restlet) and design used to implement the BioPortal REST services themselves.
To get my ConceptRestlet to accept the injection of the "ConceptServiceImpl" implementation, one first defines a setting method for that class:
public class ConceptRestlet...
private ConceptService conceptService;
public void setConceptService(ConceptService conceptService)
this.conceptService = conceptService;
}
One then specifies in the BioPortal Spring configuration file for the BioPortal Interface Layer ("applicationContext-rest.xml") the implementations to inject into ConceptRestlet. I've appended an excerpt from the configuration file:
<bean id="conceptRestlet" class="org.ncbo.stanford.view.rest.restlet.concept.ConceptRestlet" parent="abstractBaseRestlet"> <property name="conceptService"> <ref bean="conceptService" /> </property> </bean>
The above configuration specifies that "ConceptRestlet" has a setter method named "setConceptService" which accepts an injection of a bean implementation referenced by "conceptService." The following configuration (found in "applicationContext-services.xml") defines the "conceptService" bean to be "ConceptServiceImpl."
<bean id="conceptService" class="org.ncbo.stanford.service.concept.impl.ConceptServiceImpl"> <property name="ncboOntologyVersionDAO"> <ref bean="NcboOntologyVersionDAO" /> </property> <property name="ontologyFormatHandlerMap"> <ref local="ontologyFormatHandlerMap" /> </property> <property name="ontologyRetrievalHandlerMap"> <ref local="ontologyRetrievalHandlerMap" /> </property> </bean>
If a developer wanted to deploy a different "ConceptService" implementation, the person would simply need to copy the new compiled bean class file into the BioPortal deployment, change the Spring configuration file to point to this new implementation, and restart the application server.
BioPortal OntologyLoadManager Walk-Through
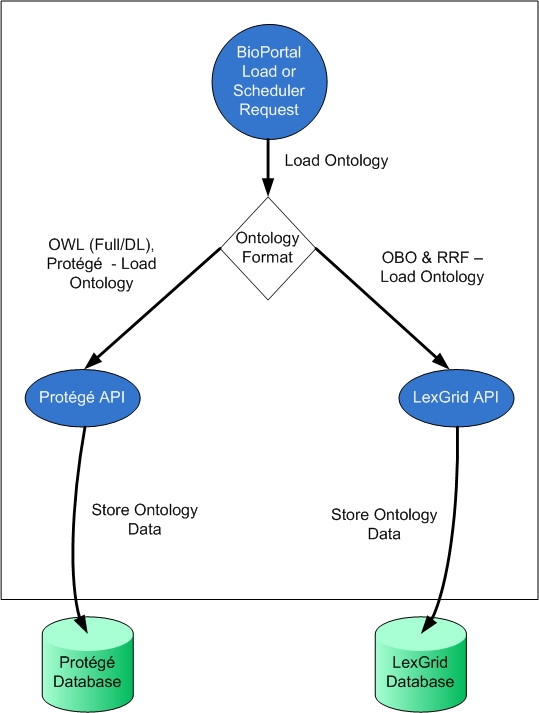
Now let's try digging into a more complex example of BioPortal Setter Injection model found in the Business Logic Layer. When new ontologies are loaded into BioPortal (either through the UI or through back end scheduled jobs), they are loaded either to two possible back-end repositories Protege or LexGrid. All OWL/Protege ontologies are loaded into Protege. All OBO/RRF (which are biomedical specific formats) are loaded into LexGrid. The following diagram presents this general architecture:
In this scenario, we have the OntologyLoadManager interface which is consumed/used by the OntologyServiceImpl class. Since the OntologyLoadManager is an interface, it has no implementation.
To get my OntologyServiceImpl to accept the injection of the OntologyLoadManager implementations, I define a setting method for that class:
class OntologyServiceImpl...
Map<String, OntologyRetrievalManager> ontologyRetrievalHandlerMap
public void setOntologyLoadHandlerMap(Map<String, OntologyLoadManager> ontologyLoadHandlerMap)
this.finder = finder;
}
One then specifies in the BioPortal Spring configuration file ("applicationContext-services.xml") the implementations to inject into OntologyServiceImpl. I've appended an excerpt from the configuration file:
<bean id="ontologyService" class="org.ncbo.stanford.service.ontology.impl.OntologyServiceImpl"> <property name="indexService"> <ref bean="indexSearchService" /> </property> <property name="ncboOntologyDAO"> <ref bean="NcboOntologyDAO" /> </property> <property name="ncboOntologyVersionDAO"> <ref bean="NcboOntologyVersionDAO" /> </property> <property name="ncboOntologyVersionMetadataDAO"> <ref bean="NcboOntologyVersionMetadataDAO" /> </property> <property name="ncboOntologyFileDAO"> <ref bean="NcboOntologyFileDAO" /> </property> <property name="ncboOntologyCategoryDAO"> <ref bean="NcboOntologyCategoryDAO" /> </property> <property name="ncboOntologyLoadQueueDAO"> <ref bean="NcboOntologyLoadQueueDAO" /> </property> <property name="ncboLCategoryDAO"> <ref bean="NcboLCategoryDAO" /> </property> <property name="ontologyFormatHandlerMap"> <ref local="ontologyFormatHandlerMap" /> </property> <property name="ontologyLoadHandlerMap"> <ref local="ontologyLoadHandlerMap" /> </property> </bean>
The above configuration specifies that "OntologyServiceImpl" has a setter method named "setOntologyLoadHandlerMap" which accepts an injection of the map named "ontologyLoadHandlerMap." This map is defined in the same Spring configuration file (applicationContext-services.xml) and references both "OntologyLoadManagerProtegeImpl" and "OntologyLoadManagerLexGridImpl" as implementations of "OntologyLoadManager." In this case, the "ontologyLoadManagerProtege" refers to "OntologyLoadManagerProtegeImpl" class and the "ontologyLoadManagerLexGrid" refers to the "OntologyLoadManagerLexGridImpl" class.
<!-- Ontology handler maps --> <util:map id="ontologyLoadHandlerMap" map-class="java.util.HashMap"> <entry> <key> <util:constant static-field = "org.ncbo.stanford.util.constants.ApplicationConstants.FORMAT_HANDLER_LEXGRID" /> </key> <ref local="ontologyLoadManagerLexGrid" /> </entry> <entry> <key> <util:constant static-field = "org.ncbo.stanford.util.constants.ApplicationConstants.FORMAT_HANDLER_PROTEGE" /> </key> <ref local="ontologyLoadManagerProtege" /> </entry> </util:map>
Here's how "OntologyLoadManagerLexGridImpl" is defined in the Spring configuration file.
<bean id="ontologyLoadManagerLexGrid" class="org.ncbo.stanford.manager.load.impl.OntologyLoadManagerLexGridImpl" parent="abstractOntologyManagerLexGrid"> </bean>
Here's how "OntologyLoadManagerProtegeImpl" is defined in the Spring configuration file.
<bean id="ontologyLoadManagerProtege" class="org.ncbo.stanford.manager.load.impl.OntologyLoadManagerProtegeImpl" parent="abstractOntologyManagerProtege"> </bean>
If a developer wanted to swap out the Protege or LexGrid loader with a different class that implemented the OntologyLoadManager interface, the person would simply need to copy the new compiled bean class file into the BioPortal deployment, change the Spring configuration file to point to this new loader class, and restart the application server.
BioPortal Format Handler Walk-Through
The BioPortal provides the ability to associate different types of ontology formats with different loaders. Now that we have walked through how the BioPortal injects two different loaders (i.e., Protege and LexGrid), this section delves into how one can associate the injected loaders with particular ontology formats.
UNDER CONSTRUCTION