OntologyRelationsMeetingNotesCompiled
Introductions
- Barry Smith: NCOR, OBO Foundry
- Alan Ruttenberg: Science Commons, OBO Foundry
- Nigam Shah: how to use relations for reasoning over biol data
- Vinay Chaudhri: SRI, knowledge building based on textbooks
- David Osumi-Sutherland: FlyBase anatomy development, modeling developmental biology
- Larry Hunter: building systems that represent large scale biol knowledge, current represent
- Peter Midford: Phenoscape, evolutionary and spatial relationships, behavior ontologies
- Melissa Haendel: anatomy and phenotype ontologies, w/ Berkeley on db querying across species
- Mike Bada: bio NLP tasks and ontologies
- Darren Natale: Protein Info Resource, protein ontology
- Karin Verspoor: comp linguist, capturing relational info in text
- Chris Mungall: phenotype/genotype representation
- Wasila Dahdul: Phenoscape, anatomy curator
- Mary Dolan: MGI
- Fabian Neuhaus: NIST
- Suzi Lewis: GO/NCBO/OBO Foundry
- David Hill: MGI ontology development and annotation
- Tanya Berardini: TAIR ontology development and annotation
Background
RO v.1.0 contained binary relations only. Relations are of 3 kinds: type-type (core RO relationships); type-instance; instance-instance
Ontologies are representations of types (the general) and the goal of the RO is to create a limited repertoire of relations linking types to build these ontologies.
For example the GO contains types only, no instances. Only something that holds for all A-s will be an assertion that holds of the type A.
'All' ontologies is not ALL ontologies, there is an understanding of what the 'scope' of all is.
ACTION ITEM: scope of 'all’ needs to be addressed taking into account that some ontologies (GO, FMA) are canonical, while others are not (for example, PRO includes mutant proteins)
Larry: what role does time play in 'all'?
Response: for process type, it plays no role, because the processes are the same regardless of time. However for continuants it does make a difference, For example, a cell membrane, isolated and experimented on, is used to draw conclusions about its role in the intact cell.
The need (and goal) is for a small suite of ontologies that describe the exceptions that can be used with the reference ontologies that represent the typical/normal.
Why keep RO small?
Some people are going to use the relations in RO in sloppy ways. The smaller the number of options the less the possibility of misuse. More per se is not bad, but if the relations are ill defined that could be bad and would poison the well.
For example, “currency has_unit $” is WRONG because there are other units of currency.
Likewise, if we have “adjacent_to” then we don't need “adjacent_to_neuron/cytoplasm/organ”. There is inherently more expressivity in the class realm than in the relation realm, thus we shall pick the former vs. the latter. But there is a counter example of a useful releation from the Cell Ontology meeting: is_expressed_on_surface_of (aka EOSO) (CD4 receptor and cell type).
The solution may be:
- Compositional relations: could be broken down into several
ontologies: RO + GO cc + cell ontology
- Mix up UI with ontologies - show the mashed up relation even if it
doesn't exist in the RO. That is: what is visible to the biologist/user is something interpretable/intuitive) and what goes into RO is whatever is necessary to capture as the formal, minimal relations)
In other words, don't just focus on one vs. the other. Further, any proposed lists of compound relational expressions should be validated by provision of definitions in terms of RO relations.
Spectrum of existing relations
One end is the English side, what appears in papers and the other side are formal definitions
GOAL is to find a middle ground: Larry: these should be driving the development of the minimal, formal ones. If 'expressed on surface of' can be broken down into existing core ones, then good - relational cross-products. In addition to the formal definitions, non-rigorous, intuitive definitions are also good.
OPEN QUESTION: expressed_in implies located_in, what relationship does expressed_in have to existing relations in RO?
Rather than creating EOSO, could also classify receptors as cell surface receptors. Describe the same biology in different ways. Instead of capturing the location in terms of the relation, capture it in the GO/CO.
all-some relations
All the instances of A stand in R to some B. For example: NOT - transforms_into, BUT YES - transformation_of. Or specifically, not every child transforms_into adult (single instance, apparently not on population scale), but every adult is a transformation of a child.
We (RO) don't want to become Tim Berners-Lee. No some-some relations.
Suzi: Are OBO_REL ids going to remain as the strings or will they transition to the numericals?
ACTION ITEM: move to numerical ids
Larry: Doesn't want type-type only for RO. Too constrained.
Barry: type-type require instance-type definitions
ACTION ITEM: Define instance level relations as well. Clarity is important. Relations need to be clarified whether they be used for type_type/type_instance/instance_instance.
Well-defined relationships are the ideal; clear to us, easy to explain to others
Chris: Should we split out part_of for instances from part_of for types? How would we do this?
- Drop type-level relations (DON"T LIKE THIS)
- Separate ids for instance and class level relations, different namespaces (THIS IS THE CHOICE, details not important right now)
- Clarify current 'punning' semantics, same id, means different thing in different contexts (however, sometimes it's hard to tell what the context is)
- not all type level relations can be defined in terms of instance level relations, need to work on both fronts
Long term proposal: Provide CommonLogic definition, port this to obo and owl as necessary - Fabian has some basic work done, Alan can help in translation but doesn't want to be the blocker. Have this in place for the next release.
Are ternary relationships possible? Yes.
Cyc solution (SRI): Rule macro predicates: Relation 'all' exists. Has_part A, B.
BFO (Basic Formal Ontology) is an upper ontology of entities, and the RO is an upper ontology of relations. The ro_bfo_bridge has domain and range constraints
e.g. id OBO_REL:has participant; the domain is span: Occurrent, and the Range is snap:Continuant
This is good in that they are separate and independent, but it is very awkward to maintain, BFO is primarily realized in owl, RO is in obo. The solution is to use a common language
ACTION ITEM: RO should have a commitment to an upper ontology and it shall be BFO.
RO tracker is located at http://sourceforge.net/tracker/?group_id=76834&atid=947684. Requests can be requirements-oriented. E.g. I need a relation for this purpose, please help me formalize it.
One solution to has_part: add the qualifiers into it: all_has_part, for example
Representing Developmental Anatomy
Presenter: David Osumi-Sutherland
Current limitations: RO can't specify timing of transitions, can't use existing part_of relations to apply to 'sometimes'. What is needed is a system for capturing stage specific part relations.
What is developmental time?
In a developmental process, many events happen in an invariant order, these events can be used as markers of developmental progress or time (standard stage series). E.g. gastrulation is a process, which can be arbitrarily divided into stages based on marker events
Stage = BFO, fiat part of process. In other words marker events: birth, death, transformation of an anatomical structure/ beginning, end, a key point in a developmental process
Timing relationships: before, simultaneous which can both be derived from before_or_simultaneous_with
(Werner joins discussion at 11:05 am local time, during David's example of Drosophila oogenesis)
Question is: How to deal with relative processes? That is, X happens during Y only during process Z?
Ternary relations? A before B during C? - but these are not supported in OWL, so have to use binary relations.
Solution: for each type for which timing is to be captured, record a reference occurrent (see slides for rest of detail). E.g. gastrulation reference_occurrent embryogenesis. There is still an issue with individual somite geneses - generalize to somitogenesis? (recurrent process) - individual timing relationships
ACTION ITEM: Add new timing relationships: begins_at_end_of, begins_during, ends_during, begins_before, ends_after, simultaneous_with, happens_during
In the future we hope to use ternary relationships. The solution for right now (that will work with the existing tools) is to use binary that can be mapped to ternary ones in the future. But David (and others) wants to reason using these relationships NOW. Argument for ternary relationship: can have multiple reference processes.
Larry's example: oocyte maturation, follicle maturation with regards to oogenesis and with regards to life of the organism.
(From Larry)
Here is my example of why a single reference occurrence doesn't work (and we need a ternary relation in order to capture the reference occurrent for a particular temporal relation):
- Oocyte fate determination <before> oocyte maturation [reference occurrent oogenesis]
- oocyte maturation <before> follicle maturation [reference occurrent lifespan]
First, note that oocyte maturation needs to have both reference occurrents in order to be able to assert these two things. These are different in that there are a bunch of oogenesis occurents (so we need the reference occurrent), but they all end before the first follicle maturation (so we need the other reference occurrent). Incidentally, we should be able to infer Oocyte fate determination <before> follicle maturation on something like the basis that oogenesis <part-of> lifespan.
ACTION ITEM: fix phrasing to timing relationships (first_exists_during vs. begin_to_exist) to be consistent.
Fabian demonstrated an example of a reasoner that uses David/Fabian's new temporal relations. Reasoning at instance level breaks, is too slow, but reasoning at type level works.
Define: what is the same and what is new? E.g. Neuroblast vs. neuron
Alan: does NOT like succeeds, doesn't like the generalization, either have specifics or general, not both.
LUNCH BREAK occurred as a way to interrupt the escalating discussion/argument.
derives_from vs. transformation_of
Presenter: Melissa Haendel
Gave specific examples of develops_from and why this needs to be transitive: heart development. But the problem is that this is not the definition of develops_from. Do we care about identity? If not, get rid of transformation_of and keep develops_from.
develops_from --derives_from --transformation_of
Darren example: A -> A-PO4, is this transformation or derivation?
Define identity function first, then can decide what is transformation and what is not. Views on what identity is, is the hard part - complicated, context-dependent. For example, consider:
A -> B -> C -> D -> E
Where B derives from A, C transformation of B, C derives from D, and E buds from D
What is the relation of A to E? Is it develops from, needed from transitive closure, good for querying
Alan's issue: if there's any lack of clarity of when to use derives_from vs. transformation_of, then the system breaks down
SOLVE THE IDENTITY PROBLEM!!!!! (and solve it now)
Cell division and budding are pretty clear: division: A -> B + C whereas budding: A -> A + B
What are the examples that are clearly transformation? Rule is to stay at same level of granularity, anatomical parts down to the cellular level (NOT molecular level) - id1 => id1
- caterpillar -> pupa -> butterfly
- neural plate transformation of neural tube
- neuron -> mature neuron
- mature leaf -> senescent leaf
- What are the examples that are clearly derives from? id1 => id2
- [mandible derives from cranial neural crest] - a bit controversial because of the 'scaffolding' involved in bone formation
- U2 neuron derives from GMC71B
- leaf primordium derives from shoot apical meristem
- neural crest derives from neural tube
Alan: Is identity reality or perspective?
If develops_from is used in cases where we don't know what to say ------ leads to a lazy way of usage.
Biologists: we KNOW it's develops from, we don't know whether it's derives from or transformation of. A process needs to have a beginning and the end, The thing that develops does not have to be there at the beginning of the process but it needs to be there at some point during the process
ACTION ITEM: make a biological definition of develops_from, test at all levels of granularity, provide cases of NOT develops_from Is transformation maturation? Is that the key? Not the single criterion. Definitely not at the RNA level. Maybe just at the organismal level. Can one have maturation independent of some process? No.
COFFEE BREAK
Spatial Relations, Coming From Rcc8
- disconnected
- externally connected - adjacent?
- equal
- partially overlapping
- tangential proper part - contained in and adjacent to
- tangential proper part inverse
- non-tangential proper part
- non-tangential proper part inverse
Don't want to make projections of 3D onto 2D.
If we want to use RCC keep the dimensions the same at the instance level. We should look at geospatial ontologies/descriptions for car (?) part manufacturing?
What is the difference between R and R' where: “town R county” and “island R' lake”? Or for a biological example: urine R bladder (contains? located?)
ACTION ITEM: have v. small meeting with spatial ontology (CAD, connectivity, etc.) experts + some biologists to adapt existing spatial relations to RO, who to invite? Tony Cohn. Thomas Bittner. Karen Pittman, Cycorp. Xenia Fiorentini, NIST. GOAL of this meeting: get spatial relations into RO
Spatial use cases
Presenter: Wasila Dahdul
teeth R bone: What is this? attached_to? connected_to? When would you use one over the other?
bona fide boundary vs. fiat boundary Does it matter? When does it matter? It is sufficient to share a boundary? Does it have to be a bona fide boundary? Connected_to is currently a fiat boundary as in the FMA: x is connected to y if X and y share a fiat (somewhat arbitrary) boundary (doesn't reflect any physical discontinuity)
Vinay: connectedness: A is connected to B means that B will move if A moves. It is grounded in physical actions/movement
Attached_to is a connected relationship where there is a large disparity between the size of the two objects. E.g. muscle attached_to bone, tooth attached_to bone, arm attached_to body, NOT body attached_to arm,
In the case of ball and socket joint they are connected via fluid. The thought experiment is whether there are more kinds of 'connected' for things, force exerted, force connectedness? fiat connectedness?
Fabian: what kind of reasoning do you want to get out of this/support with these relationship types? Both approaches have their merits. What is the purpose? What would be useful?
He suggests, just have a single relationship called 'connected' which covers both.
Barry's motion: add attachment, connected, synaptic connection, get definitions from FMA which need improvement, improve them, submit to biologists for review with examples, find out why FMA used them that way. It looks like discussions on the RO list led to use of these proposed types (with same names) in the opposite direction. Is there really confusion or not? Not.
Overlaps relationship
Peter Midford
Overlaps means two things share parts, for example: synovial joint
- A part of Bone 1
- A part_of Synovial Joint 1
- Therefore Bone 1 overlaps Synovial Joint 1
Logically, everything overlaps with its parts. Are these symmetric on the instance level but not on the type level? This seems the case for the joint example. However uterine tract overlaps urogenital system but NOT urogenital system overlaps uterine tracts
ACTION ITEM: Add symmetric_overlaps/mutally_overlaps, modify the proposed definition.
If the femur overlaps knee joint then the reverse is knee joint overlaps femur. Are both true? Is it important to capture that this is a symmetric overlap?
Wasila notes that in the spatial ontology top level contains both dependent and independent continuants: anatomical axis, anatomical axis direction, anatomical compartment, anatomical compartment boundary, anatomical gradient, anatomical region - included margins, anatomical section, anatomical surface
Character, Character State
What is a character description? Characters from a phylogenetic perspective, cover the variation in a single bone, how is it shaped differently across different species. Sample character: shape of posterior margin of lateral ethmoid has one phenotype in one species and another phenotype in another species. Of another example: the Character is the presence or absence of bone X. Or the position of bone X with regard to bone Y, where bone X may be posterior to Bone Y or Bone X may be anterior to Bone Y.
- ACTION ITEM: make GO spatial-related process definitions consistent with spatial ontology
- ACTION ITEM: add more axis-related terms to spatial list (adaxial/abaxial, apical/basical, caudal-cephalic)
- ACTION ITEM: take anything relational out of Spatial and move to RO, check out whether 'vicinity of' needs to stay
- ACTION ITEM: remove spatial stuff from FMA
- ACTION ITEM: decide what needs to stay in PATO and what needs to go into RO - punt to tomorrow
- ACTION ITEM: Define distal and proximal, ternary relationships?
- ACTION ITEM: Add medial and lateral?
Agency
Chris: is_agent.
Is there really agency ? NO because of the absence of intent. But still need something more specific than has_participant, e.g. heart process has_participant heart
Alan : NO NEED FOR AGENT! (see his very complicated slide about how one can get around this using roles and owl). The only role related relation Alan thinks is necessary: realization_of
Fabian: participates in?
Is it one or is it a bunch? punt to tomorrow
Regulates
David Hill
Fabian: can processes that regulate a process be a part of that process? Yes. Good.
Ignoring regulations of qualities for now? Yes. There are not targets in the GO that already exist.
Alan: so many things affect transcription, how do you distinguish some that regulate and some that don't
By restricting to occurrents, we avoid problems that could arise from things like 'temperature' and 'nucleotide concentration' affecting the 'rate, frequency, extent.' I.e. RNA pol II does not regulate transcription, it carries out transcription.
What about other parts of the transcription machinery? If they're always there, they do transcription. If they're sometimes there, they are regulating.
If it's static canonically, it does not regulate.
Alan has problems with the current definition: one that includes all the things based on your intuition. cite the right entities. We should have some procedure so that I can take two processes that are happening in the vicinity of the other process that let's me tell whether one is regulatory and the other is not. Right now, it's not workable. Define rate. Who are the participants? What is the rate of a process? Duration makes sense. What is frequency of a process? His sense is that the current definition is completely lexically driven. Define in terms of a continuant which feeds into the process (did I catch this right?). In as many processes as possible.
There are participants in the BMP signaling pathway that directly do the regulation of transcription. There are proximate processes that affect transcription.
Suzi: it's the other way around. There is big effort in GO to use regular syntax when naming things. These things got named this because of what they did.
You can't change a canonically static process
The rate of a process is how many times does it get initiated? ADDED POST MEETING: The rate is how long the process takes form start to finish. The frequency is how often it gets initiated. The extent is how far along it has gone from start to finish. (DPH)
Larry: Can Alan improve 'regulates' any further?
Qualities of occurrents? Can they exist? ADDED POST MEETING: Currently there are PATO terms that describe changes in qualities of occurrents such as id: PATO:0000161 name: rate subset: attribute_slim is_a: PATO:0001239 ! monadic quality of occurrent
Causation - regulation - How far back do you go? What is the scope? Basic metabolic processes regulated EVERYTHING. Where is the beginning and end? What is direct and indirect?
possible solution: use comparatives, why this instead of that? The comparative approach helps us do science. All of experimentation is comparative.
Why isn't the production of BMP part of this process? Probably yes, but this figure was taken out of a paper.
Is regulates transitive? If A regulates B and B regulates C, does A regulate C?
ACTION ITEM: improve on the current 'regulates' definition referring to continuants which interact and fulfill their functions, then see if this definition works and can be applied to the use case/s that we have
When known, the most proximal step to a process, should be the regulator of that process. In the example, binding of SMAD4 complex regulates transcription, all the previous stuff regulates transcription by inference.
Do we need 'regulates' AND 'indirectly_regulates'? Not really.
Is regulation a mix of instructive and permissive signals?
Hh signaling regulates limb development? Melissa/David: yes David OS: BS! -lots of argument, David H changes his mind, Hh signalling part of limb development. In summary, instructive signals would be part of the process
BMP signaling is required for dorsal/ventral patterning, it is instructive, it needs to be a part of.
Darren: If all we know that BMP binding the receptor results in change in tsc, then create BMP signaling pathway regulates transcription. As we know more, add more information to the ontology, change the annotation to something more specific.
Larry: why take away the more general and only keep the detail?
How many senses of 'regulates' are being lumped together in the relationship 'regulates'?
END DAY ONE - GOING TO DINNER at 6:20 pm after beginning at 8:50 am
Day 2: starting at 9:13 am with tasks from last night
Develops_from homework
Biologist, logicians to try to come up with definitions of develops_from, discuss need for transformation_of, derives_from.
Biological
Melissa/David: Biological definition of develops_from (at the cellular level) and the continuity of cell lineage.
Darren: c develops from d iff there is a continuity of lineage extending from d to c, where:
- the existence of d preceds the existence of C
- some portion of the material of d is retained in the material of c
- one can trace the event or set of events linking d to c
What is 'material'?
Alan: still need and should have developmental specific definition, build the specific definitions (developmental, protein) first, then build the general one later
- cell1 parent_of cell 2 (single division process occurs)
- cell1 in_lineage_with cell2, if cell1 = cell2 or cell2 is some sequence of parent relationships to the cell2
(develops_from within a single organism) - gets around Larry's lung developed from his grandmother's ovary
anatomical structure 1 develops from anatomical structure 2 iff some cell in anatomical structure 2 in lineage with some cell in anatomical structure 1.
Is there a problem with immaterial entities? Eg. blastocoel. Not really, we don't care about them. What about secretions, things that are built out of secreted structures? How do we extend to those? Use 'secretes'?)
This basic definition appears to apply on multiple levels: cellular, biochemical, anatomical structures and organisms. We will try this out.
Suzi: some events that result in something new, some events that result in 'rearrangement of material'
Alan: we want at least the cell biological level defined, good logical definition in the terms that biologists use and understand, build from the cellular level up.
Logical
(2) ontologist's justification of why the two children terms are important ;
Biologists don't care explicitly, use it implicitly. Ignore the two children for now.
Homology
Melissa: homology relation? 'descends_from' homologous structures vs. analogous structures. For example, mouse limb homologous to human limb, but bird wing is analogous to bat wing character = quality (shape of bone margin) character state = variability in quality Peter: Review of tree building based on three characters, looking at the various character states independently reveals different things about the individual ancestral states. Comparing two trees derived from two different sets of characters reveals different phylogenetic relationships. Homology evidence codes: (in use by Phenoscape using Phenote)
- inferred from morphological similarity
- inferred from positional similarity
- inferred from developmental similarity
- inferred from compositional similarity
- inferred from gene expression similarity
- inferred from phylogeny
Definition: (see RO meeting wiki but remove taxon reference and replace by 'population', remove the Instance 3-ary relation - Melissa editing now)
directly_descends_from = (definition, get rid of partially a copy, get rid of determination is instance_level
descends_from = ? in_lineage_with from the earlier discussion on develops_from
This looks good to Barry, fix italicization issue with definition
common ancestor forelimb
/ \
/ \ descends_from
/ \
/ \
bat wing human arm
bat wing <-homologous_to -> human arm
ACTION ITEM: Two relations: 'descends_from' (not the final string), homologous_to both with definitions - generalize definition, work on last bit of definition relating to genetic material and genetic programs
What do you use to describe the relationship between a parent anatomical structure and a child anatomical structure
Darren: current definition is restricted to anatomical structures, huge problem for proteins/genes
Chris: structure of definition is fine, need to generalize homology to apply to proteins/genes, anatomical structure could broadly refer to character states
Melissa: need homologous_to NOW for anatomical structures
Note: Just because two anatomical structures are homologous, it does not mean that its parts are homologous. Structures that are homologous could use completely different programs.
David OS: transitive closure is important only between two 'generations'
- 1001
- 0001
- 0101
- 0110
- 0100
David OS: evolved_from relationship. The homologous_to relationship does not involve time, can hold between simultaneously existing entities: human arm and bat wing. In contrast, evolves_from does involve time: human arm and 'ancestral forelimb'
Is evolves_from built from directly_descends_from? Yes. Figure out strings to make them parallel.
BREAKTIME
Regulates discussion: not reopened to avoid non-constructive and never ending discussion.
Action Item: David, Tanya and Chris will come up with a more rigorous definition of 'regulates' and present that to the group for review.
Epistemic relations or how do we know what we know?
Presenter: Larry Hunter
- supports
- is_supported_by (inverse)
- contradicts
- is_contradicted_by (inverse)
What is scientific evidence? (evidence that supports a hypothesis - 'citeable'?)
- peer review
- capable of being communicated
- secret information can't be scientific evidence
Making physical objects scientific evidence is a problem???
- "bug" with pins in it? scientific specimen: is this evidence?
- how about clones?
information entity - something that is communicated
GO gene_association file (GAF) use case
3 epistemic columns:
- DB:reference: <assertion> is_supported_by <DB:reference>
- NOT qualifier - flips evidence, not assertion: <assertion> is_contradicted_by <DB:reference>
- evidence code
How to handle one piece of evidence that supports two (or more) assertions in different ways. I.e. sentence 1 provides info for IDA and IC, how do we capture these multiple roles?
- sub-relations: subclass binary supports, e.g. supports_by_direct_assay
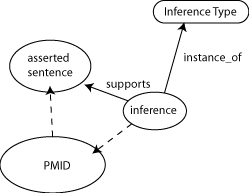
- ternary relations: Make supports a ternary relation: supports <assertion> <evidence> <evidence type>
- PMID = ECO type
- represent the inference
- asserted sentences have a speaker
- asserted sentences = things we believe are true
- (When all fails, speak louder and/or faster.)
- partial constraints for supports?
- Barry: add to ROIL
TAIR Relations
Tanya: has(function)
ACTION ITEM: table: map relationship_types to RO relations if known, if not put?, also add range of things that go into term2, send to group (obo-discuss, ro). For protein modification, use psi-mod (potential target for)
Biochemical relations
Presenter: Mike Bada
Built GO to CheBI cross products
Functions defined in terms of processes by which they can be realized:
kinase function = catalytic function THAT is_realized_in GO:kinase activity
Definitions are wrong!
THEY WANT: Catalytic processes and Catalytic activities defined in terms of processes they are catalyzing:
GO:kinase activity = GO:molecular_function THAT results_in_catalysis_of GO:phosphorylation
This (Mike thinks) necessitates an ontology of types of (bio)chemical reactions. GO lays out potential to do something vs. the doing something. These both exist. We will not instantiate BOTH function AND functioning. It is simpler to make links between functions and processes.
Process(function) = realization of function, and Function (process) = realized_as process
Alan: automatic generation of one term when the other is made
DavidH: how about creating links between MF and BP and the relationship describes that link?
Discussion about Mike's definitions and how they could be improved.
David H: We will create a relationship describing that a gene product has the potential to execute a molecular function (has disposition to realize?). This way the MF ontology describes the realization (execution) of a molecular function. These executions will then be allowed to be part_of BPs. All MFs are part_of some type of BP.
TMRs, cross-products
ACTION ITEM: Larry will create and use too many relations (TMRs) Mike has made now as 'macros' or shortcuts while the RO people write the underlying framework that will support them. If some of them don't break down properly, these will be fixed. Relations will be sent to Alan together with Mike for training. Code named: Mike And Chris Relations Ontology, or ROC, ROIL, MAC-RO, MAC-ROP
Chris: cross products, GO biological process and cell ontology
There are a bunch of different proposed relations: see ro_proposed: aka Chris' macros (how many? 12, development, anatomy and cell, for chemicals about 10)
ACTION ITEM: find an example of Chris' macro where there are two relations that identify two participants to verify claim - source and destination for a transport event
ACTION ITEM: again, find a way to translate the macros into a formal definition
Pattern: x results in A x2; A is term from another ontology
Alternate: for some y A(y); participant (x2,y) ^
(tried to transcribe the above from Fabian's writing on the board but missed some parts)
Necessary is easy, necessary and sufficient is hard
Summary of Action Items
- Scope of 'all’ needs to be addressed taking into account that some ontologies (GO, FMA) are canonical, while others are not (for example, PRO includes mutant proteins)
- Move to numerical ids
- Define instance level relations as well. Clarity is important. Relations need to be clarified whether they be used for type_type/type_instance/instance_instance.
- RO should have a commitment to an upper ontology and it shall be BFO.
- Add new timing relationships: begins_at_end_of, begins_during, ends_during, begins_before, ends_after, simultaneous_with, happens_during
- Stages and temporal relations: are there cases where we do not need the reference process?
- Fix phrasing to timing relationships (first_exists_during vs. begin_to_exist) to be consistent.
- Make a biological definition of develops_from, test at all levels of granularity, provide cases of NOT develops_from
- Have v. small meeting with spatial ontology (CAD, connectivity, etc.) experts + some biologists to adapt existing spatial relations to RO, who to invite? Tony Cohn. Thomas Bittner. Karen Pittman, Cycorp. Xenia Fiorentini, NIST. GOAL of this meeting: get spatial relations into RO
- Spatial relations: add to ro attachment, connected, synaptic (use fma definitions, work on them, circulate to biologists to vet)
- Add symmetric_overlaps/mutally_overlaps, modify the proposed definition.
- Make GO spatial-related process definitions consistent with spatial ontology. Fix, harmonize, extract from appropriate places -- anything relational from spatial.obo to ro
- Add more axis-related terms to spatial list (adaxial/abaxial, apical/basical, caudal-cephalic). Make precise what is meant by medial, lateral -- dorsal, ventral -- proximal, distal -- anterior, posterior
- Take anything relational out of Spatial and move to RO, check out whether 'vicinity of' needs to stay
- Remove spatial stuff from FMA
- Decide what needs to stay in PATO and what needs to go into RO - punt to tomorrow
- Define distal and proximal, ternary relationships?
- Add medial and lateral?
- Improve on the current 'regulates' definition referring to continuants which interact and fulfill their functions, then see if this definition works and can be applied to the use case/s that we have
- Two relations: 'descends_from' (not the final string), homologous_to both with definitions - generalize definition, work on last bit of definition relating to genetic material and genetic programs. new relations descends_from (already used so need a different string!), homologous_to can be inferred.
- Evidence: add supports and contradicts relationships, small working group for domain and range
- David, Tanya and Chris will come up with a more rigorous definition of 'regulates' and present that to the group for review.
- Table: map relationship_types to RO relations if known, if not put? Also add range of things that go into term2, send to group (obo-discuss, ro). For protein modification, use psi-mod (potential target for)
- Tair: make table of tair relations, column for ro term if it exists, send to ro group make request for needed terms
- Larry will create and use too many relations (TMRs) Mike has made now as 'macros' or shortcuts while the RO people write the underlying framework that will support them. If some of them don't break down properly, these will be fixed. Relations will be sent to Alan together with Mike for training. Code named: Mike And Chris Relations Ontology, or ROC, ROIL, MAC-RO, MAC-ROP
- Find an example of Chris' macro where there are two relations that identify two participants to verify claim - source and destination for a transport event
- Again, find a way to translate the macros into a formal definition